-Search query
-Search result
Showing all 46 items for (author: lu & df)

EMDB-35369: 
Cryo-EM structure of RBD/E77-Fab complex
Method: single particle / : Lu DF, Zhang ZC

EMDB-27701: 
Focused map (monomer A) for Arabidopsis SPY in complex with GDP-fucose
Method: single particle / : Kumar S, Zhou Y, Dillard L, Borgnia MJ, Bartesaghi A, Zhou P

EMDB-27702: 
Focused map (monomer B) for Arabidopsis SPY in complex with GDP-fucose
Method: single particle / : Kumar S, Zhou Y, Lucas D, Borgnia MJ, Bartesaghi A, Zhou P
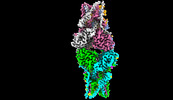
EMDB-35503: 
Cryo-EM structure of Stimulator of interferon genes
Method: single particle / : Lu DF, Shang GJ

EMDB-35504: 
Structure of Stimulator of interferon genes/ligand complex
Method: single particle / : Lu DF, Shang GJ

EMDB-28776: 
Structure of VSD4-NaV1.7-NaVPas channel chimera bound to the hybrid inhibitor GNE-1305
Method: single particle / : Kschonsak M, Jao CC, Arthur CP, Rohou AL, Bergeron P, Ortwine D, McKerall SJ, Hackos DH, Deng L, Chen J, Sutherlin D, Dragovich PS, Volgraf M, Wright MR, Payandeh J, Ciferri C, Tellis JC

EMDB-28777: 
Structure of VSD4-NaV1.7-NaVPas channel chimera bound to the acylsulfonamide inhibitor GDC-0310
Method: single particle / : Kschonsak M, Jao CC, Arthur CP, Rohou AL, Bergeron P, Ortwine D, McKerall SJ, Hackos DH, Deng L, Chen J, Sutherlin D, Dragovich PS, Volgraf M, Wright MR, Payandeh J, Ciferri C, Tellis JC

EMDB-28778: 
Structure of VSD4-NaV1.7-NaVPas channel chimera bound to the arylsulfonamide inhibitor GNE-3565
Method: single particle / : Kschonsak M, Jao CC, Arthur CP, Rohou AL, Bergeron P, Ortwine D, McKerall SJ, Hackos DH, Deng L, Chen J, Sutherlin D, Dragovich PS, Volgraf M, Wright MR, Payandeh J, Ciferri C, Tellis JC

EMDB-28779: 
Structure of VSD4-NaV1.7-NaVPas channel chimera bound to the hybrid inhibitor GNE-9296
Method: single particle / : Kschonsak M, Jao CC, Arthur CP, Rohou AL, Bergeron P, Ortwine D, McKerall SJ, Hackos DH, Deng L, Chen J, Sutherlin D, Dragovich PS, Volgraf M, Wright MR, Payandeh J, Ciferri C, Tellis JC

EMDB-27177: 
sd1.040 Fab in complex with SARS-CoV-2 Spike 2P glycoprotein
Method: single particle / : Abernathy ME, Barnes CO

EMDB-15905: 
Cryo-EM structure of the E.coli 70S ribosome in complex with the antibiotic Myxovalargin B.
Method: single particle / : Koller TO, Graf M, Wilson DN

EMDB-14121: 
Cryo-EM structure of the E.coli 50S ribosomal subunit in complex with the antibiotic Myxovalargin A.
Method: single particle / : Koller TO, Beckert B, Wilson DN

EMDB-29002: 
Nucleocapsid monomer structure from SARS-CoV-2
Method: single particle / : Casasanta M, Jonaid GM, Kaylor L, Luqiu W, DiCecco L, Solares M, Berry S, Kelly DF

EMDB-29072: 
SARS-CoV-2 Nucleocapsid dimer structure determined from COVID-19 patients
Method: single particle / : Casasanta M, Jonaid GM, Kaylor L, Luqiu W, DiCecco L, Solares M, Berry S, Kelly DF

EMDB-28189: 
SARS-CoV-2 Spike in complex with biparatopic nanobody BP10
Method: single particle / : Pymm PG, Glukhova A, Tham WH
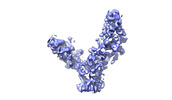
EMDB-28190: 
SARS-CoV-2 RBD in complex with biparatopic nanobody BP10 local refinement
Method: single particle / : Pymm PG, Glukhova A, Tham WH

EMDB-28816: 
Wild type P53 dimer structure from human cancer cells
Method: single particle / : Solares M, Kelly DF

EMDB-28817: 
P53 monomer structure
Method: single particle / : Solares M, Kelly DF
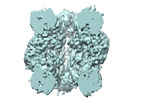
EMDB-27654: 
A subtomogram average of H. neapolitanus Rubisco within alpha-carboxysomes
Method: subtomogram averaging / : Metskas LA, Blikstad C, Laughlin T, Savage DF, Jensen GJ

EMDB-31232: 
Structural basis for the tethered peptide activation of adhesion GPCRs
Method: single particle / : Ping YQ, Xiao P, Yang F, Zhao RJ, Guo SC, Yan X, Wu X, Sun JP

EMDB-31254: 
GPR114-Gs-scFv16 complex
Method: single particle / : Ping Y

EMDB-24002: 
Cryo-EM map of the FLP cargo within the T. maritima encapsulin
Method: single particle / : LaFrance BJ, Savage DF, Nogales E

EMDB-24001: 
Thermotoga maritima encapsulin shell
Method: single particle / : LaFrance BJ, Nogales E, Savage DF

EMDB-22518: 
SpCas9 delta4CE Ternary Complex
Method: single particle / : Sham A, Laughlin TG, Savage DF

EMDB-24147: 
Nucleocapsid protein from the SARS-CoV-2 Coronavirus
Method: single particle / : Casasanta MA, Kelly DF

EMDB-23566: 
The SARS-CoV-2 spike protein receptor binding domain bound to neutralizing nanobodies WNb 2 and WNb 10
Method: single particle / : Pymm P, Glukhova A, Tham WH

EMDB-23567: 
Cryo-EM map of SARS-CoV-2 Spike protein bound to neutralising nanobodies WNb2 and WNb10 (C3 symmetry)
Method: single particle / : Pymm P, Tham WH, Glukhova A

EMDB-23568: 
Cryo-EM map of SARS-CoV-2 Spike protein in complex with neutralising nanobodies WNb2 and WNb10 (C1 symmetry)
Method: single particle / : Pymm P, Tham WH, Glukhova A

PDB-7lx5: 
The SARS-CoV-2 spike protein receptor binding domain bound to neutralizing nanobodies WNb 2 and WNb 10
Method: single particle / : Pymm P, Glukhova A, Tham WH

EMDB-22982: 
Nucleocapsid protein from the SARS-CoV-2 coronavirus
Method: single particle / : Casasanta MA, Kelly DF

EMDB-22827: 
P53 tetramer from Glioblastoma
Method: single particle / : Solares MJ, Kelly DF

EMDB-22124: 
SARS-CoV-2 S 2P negative-stain EM
Method: single particle / : Barnes CO, Bjorkman PJ

EMDB-22125: 
SARS-CoV-2 S 2P trimer complexed with COV57 polyclonal Fabs
Method: single particle / : Barnes CO, Bjorkman PJ

EMDB-22126: 
SARS-CoV-2 S 2P trimer complexed with polyclonal Fabs from recovered COVID-19 individual COV21
Method: single particle / : Barnes CO, Bjorkman PJ

EMDB-22127: 
Structure of the SARS-CoV-2 spike glycoprotein in complex with the C105 neutralizing antibody Fab fragment (state 1)
Method: single particle / : Barnes CO, Bjorkman PJ

EMDB-22128: 
Structure of the SARS-CoV-2 spike glycoprotein in complex with the C105 neutralizing antibody Fab fragment (state 2)
Method: single particle / : Barnes CO, Bjorkman PJ

EMDB-22094: 
CryoEM structure of the holo-SrpI encapsulin complex from Synechococcus elongatus PCC 7942
Method: single particle / : LaFrance BJ, Nichols RJ, Phillips NR, Oltrogge LM, Valentin-Alvarado LE, Bischoff AJ, Savage DF, Nogales E

EMDB-22095: 
CryoEM structure of the apo-SrpI encapasulin complex from Synechococcus elongatus PCC 7942
Method: single particle / : LaFrance BJ, Nichols RJ, Phillips NR, Oltrogge LM, Valentin-Alvarado LE, Bischoff AJ, Savage DF, Nogales E

PDB-6x8m: 
CryoEM structure of the holo-SrpI encapsulin complex from Synechococcus elongatus PCC 7942
Method: single particle / : LaFrance BJ, Nichols RJ, Phillips NR, Oltrogge LM, Valentin-Alvarado LE, Bischoff AJ, Savage DF, Nogales E

PDB-6x8t: 
CryoEM structure of the apo-SrpI encapasulin complex from Synechococcus elongatus PCC 7942
Method: single particle / : LaFrance BJ, Nichols RJ, Phillips NR, Oltrogge LM, Valentin-Alvarado LE, Bischoff AJ, Savage DF, Nogales E

EMDB-0378: 
p53 dimer assembly
Method: single particle / : Kelly DF, Dearnaley WJ, Varano AC

EMDB-1807: 
Saccharomyces cerevisiae ribonucleotide reductase hole complex at the presence of dATP
Method: single particle / : Fairman JW, Wijerathna SR, Ahmad MF, Xu H, Nakano R, Jha S, Prendergast J, Welin RM, Flodin S, Roos A, Nordlund P, Li Z, Walz T, Dealwis CG

EMDB-1877: 
CryoEM structure of the chromatin remodelling factor ISW1a bound to a mononucleosome (45N29)
Method: single particle / : Frouws TD, Richmond TJ

EMDB-1878: 
CryoEM structure of the remodelling factor ISW1a bound to a mononucleosome (45N0)
Method: single particle / : Frouws TD, Richmond TJ

EMDB-1409: 
The EM structure of human DNA polymerase gamma reveals a localized contact between the catalytic and accessory subunits.
Method: single particle / : Yakubovskaya E, Lukin M, Chen Z, Berriman J, Wall JS, Kobayashi R, Kisker C, Bogenhagen DF

EMDB-1410: 
The EM structure of human DNA polymerase gamma reveals a localized contact between the catalytic and accessory subunits.
Method: single particle / : Yakubovskaya E, Lukin M, Chen Z, Berriman J, Wall JS, Kobayashi R, Kisker C, Bogenhagen DF
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
